China, the US should both be on board to drive real climate action: director of Climate Group

Editor's Note:
With the conclusion of COP28, or the 28th session of the Conference of the Parties to the UN Framework Convention on Climate Change on December 12, 2023, a "historic" climate deal was inked, which, for the first time, pledged to transition away from the use of fossil fuels while boosting renewable energy. Representatives from nearly 200 countries agreed, at the summit, to begin reducing the global use of fossil fuels, drawing worldwide attention. At the summit, China talked with every relevant party to find an acceptable solution to promote the success of the COP28. Over the years, the efforts that China has made in climate change have won wide recognition and the cooperation between China and the US is also of importance to the world. On the heels of COP28, Global Times reporter Xie Wenting (GT) spoke with Champa Patel (Patel), Executive Director for Governments and Policy at the Climate Group, an NGO dedicated to climate change, on issues related to COP28 and global cooperation on climate change among other topics.
GT: What are your thoughts on the outcomes from COP28 and what are the key takeaways from conference? How do you believe this conference has contributed to global climate action?
Patel: The main takeaway from COP28 was the first explicit recognition that the world has to transition away from fossil fuels. While it did not go as far as saying "phase-out" from fossil fuels, this still sends a strong signal on what is expected of countries and that a fossil-fuel-free future is the only way forward. It sends an important message that fossil fuels are on their way out, and might not be worth investing in.
With that in mind, it was great to see commitments on tripling renewables and doubling down on energy efficiency, measures which earlier on in the year had been taken on by the G20 in its communique as well. There was also a welcome recognition that nation states should work closely with subnational governments - as the level of government often closest to impacted communities - to set climate action plans and ensure an integrated multi-level approach.
But there were significant gaps as well. Climate finance was not the focus as much as it should have been.
In many ways, we do not have a crisis of ambition - most countries are signed up to what needs to be done to achieve net zero - but for many developing economies this will require substantial investment and funding. Where will the money from? There is still much more that needs to be negotiated on new sources of climate finance and how existing funds can be scaled up - so there, we do see a crisis of ambition.
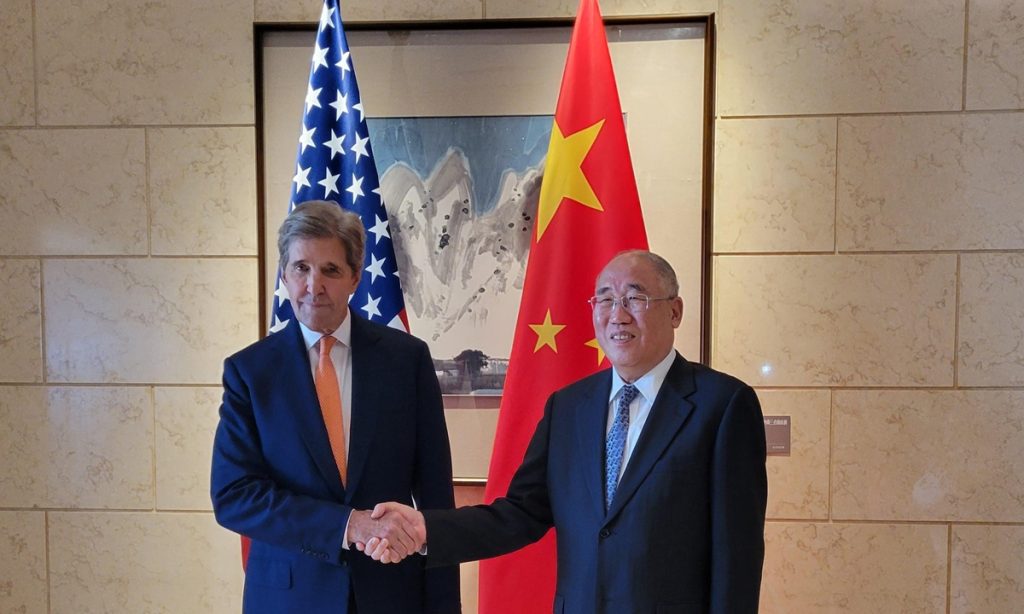
GT: The China-US climate cooperation has been a significant topic of discussion in recent years. In your opinion, what are the key areas in which China and the US can collaborate effectively to address climate change? How can this cooperation be strengthened further?
Patel: Prior to COP28, China and the US released "The Sunnylands Statement on Enhancing Cooperation to Address the Climate Crisis." The two sides have also agreed to establish a working group on enhancing climate action in the 2020s, to advance discussions on methane, the energy transition, and resource efficiency among others. This provides an important vehicle for enhanced cooperation. Interestingly, and for Climate Group more importantly, was an explicit recognition of the role of subnational cooperation bringing together states, regions, and cities in climate action. This is really important as local governments are often best placed to know the specific needs of their communities.
To drive real climate action, we need to have both China and the US on board - without them, action is meaningless. So it's great to see climate as one of the few areas which is not prey to the great power competition. The climate crisis has the opportunity to bring the great powers together. We need China and the US on board, not just because of political power or their large economies, but also because they are facing the devastating impacts of climate change within their own countries, whether heatwaves, flooding, or droughts. So these steps are a positive sign but much more needs to be done to drive action further and faster as time is critical to ward against a 1.5-degree rise in temperature.
GT: How can NGOs contribute toward fostering collaboration and driving impactful change? In light of recent COP28 commitments, what specific actions or policies do you believe the US and China should prioritize to accelerate their transition to a low-carbon economy?
Patel: NGOs have an important role to play as they can support and foster partnerships, encourage peer exchange and facilitate relationship building between China and the US. They can also help identify policy measures that can accelerate climate impact.
Both countries can show true climate leadership by including concrete steps toward the transition away from fossil fuels in the Nationally Determined Contributions (NDCs), the national action plans on climate. That would send an incredibly strong signal to the rest of the world.
GT: Looking ahead, what are your expectations for future China-US climate cooperation? How can this partnership evolve and expand to tackle emerging challenges and seize new opportunities in the fight against climate change?
Patel: Looking ahead, it is critical that the US and China identify concrete projects, initiatives, and funding that can help accelerate climate action. There is an opportunity to drive leadership not just from their respective countries but also to model what is needed from other major powers and developed economies.
The signal on action needed on methane sent through "The Sunnylands Statement on Enhancing Cooperation to Address the Climate Crisis" is important. It is the first time China has mentioned methane, as it focuses its mind on this short-lived pollutant that is often sidelined when considering decarbonization measures. But tackling methane emissions is essential as, arguably, we cannot stay within 1.5 degrees of temperature rise without also addressing methane emissions.
By coming together, the two countries can help unlock global ambition and provide a model of leadership that is sorely needed to drive forward faster climate action.